Harmonization Guides & Manuals
Installing and using Armadillo and DataSHIELD
Analysis infrastructure protocols
EU Child Cohort Governance Document, last updated in December 2019
EU Child Cohort Variable Catalogue: Guide for cohort data managers to upload source variables and harmonizations
LifeCycle protocol for harmonization and quality checks of core variables (WP1), last updated in May 2020
LifeCycle protocol for harmonization and quality checks of WP3.1.2: Ethnicity (WP3), last updated in May 2020
LifeCycle protocol for harmonization and quality checksof WP3.1.4: Lifestyle (WP3), last updated in May 2020
LifeCycle protocol for harmonisation and quality checks of WP3.3.3: Urban Environment Stressors (WP3), last updated in June 2021
LifeCycle protocol for harmonization and quality checks of cardio-metabolic health variables (WP4), last updated in April 2020
LifeCycle protocol for harmonization and quality checks of respiratory health variables (WP5), last updated in April 2020
LifeCycle protocol for harmonization and quality checks of mental health variables (WP6), last updated in April 2020
LifeCycle protocol for harmonization and quality checks of Neuroimaging data (WP6), last updated in Sept 2022
LifeCycle protocol for harmonization and quality checks of methylation data in an ExpressionSet (WP8), last updated in Sept 2022
- ATHLETE protocols for additional harmonizations on chemical exposome, fetal growth and placenta function, mental cardiometabolic and respiratory outcomes
Variable Catalogue
The variable Catalogue of the EU Child Cohort Network can be found here: Variable Catalogue (molgeniscloud.org)
As part of the LifeCycle project, the EU Child Cohort Network Variable Catalogue has been developed. The aim of the catalogue is to provide an overview of available data in the EU Child Cohort Network in order to facilitate long-term collaborations between existing pregnancy and child cohorts. The catalogue was built using the MOLGENIS software platform for scientific data (https://molgenis.org, Swertz et al., Bioinformatics, 2014). The catalogue is an open and accessible online platform that documents what type of harmonized variables (LifeCycle variables) are available from each participating cohort. It also documents how each cohort has harmonized these variables, including the raw variables (source variables) used by the cohorts, which were used to derive the LifeCycle variables. No actual data are contained in the online catalogue. The catalogue also contains a description of each cohort included in the LifeCycle project with links to the codebooks.
Central Analysis Server
The Central Analysis Server can be found here: https://lifecycle.analysis.molgenis.org
We have implemented a data management and analysis platform based on DataSHIELD (www.datashield.org), and a metadata catalogue describing the data made available based on MOLGENIS (http://molgenis.org) within a governance framework in which responsibility for data management and security is maintained by the source studies.
We have chosen a central analysis server (central setup) of R-Studio to facilitate a central R-environment to do DataSHIELD analyses on. The reason we have set up the infrastructure this way is because of security, reproducibility of research environments, a way to share analysis easily, and to have some version of control on the analysis-scripts.
Security
We rather do not expose servers which contain data directly to the internet. The DataSHIELD network is now secured in 2 layers. One is the firewall between the Armadillo or Opal instances and the analyses server and the other one is between the central analysis server and the different research-facilities where research perform there analysis. Everyone who wants access to the central analysis server has to do a request for opening up the firewall to their facility or specific workstation (if needed).
Reproducibility
Because of the wide availability of R-packages and other tooling, we wanted to have a shared analysis environment. The idea is that all work spaces in the central-analysis-server are the same. If researchers want to use a new R-package it should be installed globally so everyone can work with it.
Sharing analysis and version control
R-Studio is fully integrated with Git (version control) and a central place where we can work on our meta analysis. In Git you are able to work with each other on one or more research projects.
This is, in short, how we have setup the environment and what choices were made to facilitate the researchers as much as possible.
Statistical methods database
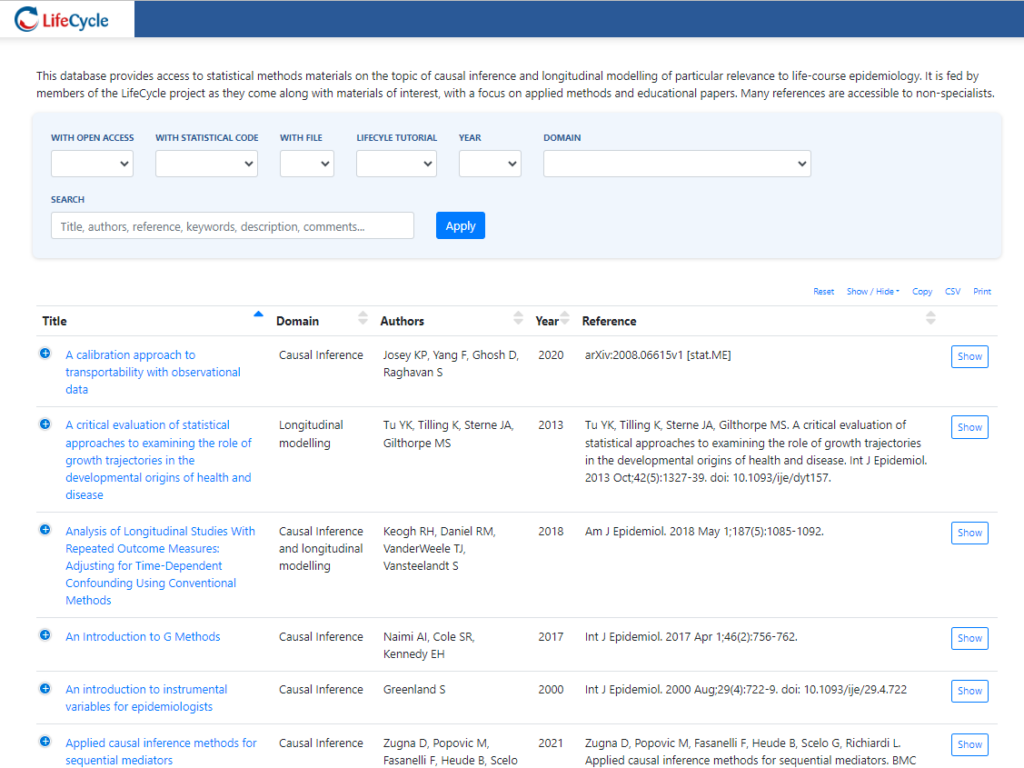
Please find the statistical methods database here: https://lifecycle.cpo.it/
The statistical methods database provides access to statistical methods materials on the topic of causal inference and longitudinal modelling of particular relevance to life-course epidemiology. It is fed by members of the LifeCycle project as they come along with materials of interest, with a focus on applied methods and educational papers. Many references are accessible to non-specialists. The website can also be accessed via the Variable Catalogue.